Table of contents
Open Table of contents
Know Your RAG
This notebook demonstrates an end-to-end workflow for generating a domain-specific evaluation dataset and using it to benchmark two RAG architectures on epidemiological literature. We apply the methodology from Know Your RAG (Lima et al., 2024) to articles from the Preventing Chronic Disease (PCD) journal — a CDC open-access publication focused on public health research — and compare an AgenticRAG system against a NaiveRAG baseline using RAGAS metrics.
Retrieval-Augmented Generation (RAG) has emerged as the dominant pattern for building question-answering systems over domain-specific corpora. Yet evaluating RAG in specialized domains — where factual precision matters and hallucinations can have real consequences — remains a hard, open problem.
Why does this matter for public health? RAG systems built for public health applications — clinical decision support, disease surveillance, policy guidance — must be rigorously tested. A system that retrieves the wrong evidence or generates a plausible-sounding but incorrect answer about chronic disease risk factors is not just unhelpful: it can actively mislead decisions about population health. Systematic evaluation with realistic, domain-grounded questions is a prerequisite for deploying these systems responsibly.
What you will learn:
- How to load and chunk scientific HTML articles from the CDC corpus
- How to generate section-aware evaluation questions (factual and reasoning) using LLMs
- How to filter questions using LLM-based critique metrics
- How to index a corpus and evaluate two RAG systems (Agentic and Naive) using RAGAS
- How to compare results in a structured evaluation table
Before you begin: All custom modules used in this notebook (
loader.py,rag_agent.py,rag_rerank.py,vectorstore.py) are available at github.com/mayerantoine/know-your-rag. Clone the repo and install dependencies before running any cells below.
Notebook Overview
This notebook is organized into four main parts:
Part 1 — Data Preparation
Load PCD articles from the CDC open dataset, parse their HTML structure, and extract semantically meaningful chunks aligned with paper sections (Abstract, Methods, Results, Discussion, etc.).
Part 2 — Evaluation Dataset Generation
Apply the Know Your RAG taxonomy to generate two types of evaluation questions per chunk:
- Fact-single questions (Results / Methods sections): single-fact, directly answerable from the text
- Reasoning questions (Discussion / Abstract / Conclusion sections): require inference across multiple statements
Each question goes through an LLM-based critique pipeline that scores groundedness, feasibility, standalone quality, and usefulness — filtering out low-quality questions before they reach the evaluation set.
Part 3 — RAG System Evaluation
Index the corpus into a ChromaDB vector store and evaluate two RAG architectures:
- AgenticRAG: multi-agent loop (search → evidence scoring → answer synthesis) using the OpenAI Agents SDK
- NaiveRAG: single-pass semantic retrieval → ColBERT reranking → LLM generation
Evaluation uses RAGAS metrics: AnswerCorrectness and Faithfulness.
Part 4 — Comparison Side-by-side mean score table comparing both RAG systems on the generated evaluation dataset.
import os
import pandas as pd
from sodapy import Socrata
import pathlib
from pathlib import Path
from typing import List, Dict, Any, Optional, Union, Tuple
from langchain_huggingface import HuggingFaceEmbeddings as lgHuggingFaceEmbeddings
from loader import download_file,get_data_directory,extract_zip_files,load_html_files
from dotenv import load_dotenv
from pydantic import BaseModel, Field
import openai
from ollama import chat
from rag_agent import AgentConfig,AgenticRAGPart 1 — Data Preparation
1.1 Dataset: CDC Text Corpora for Learners
We use the CDC Text Corpora for Learners dataset, available on data.cdc.gov. This dataset provides full-text HTML articles from three CDC publications:
- MMWR (Morbidity and Mortality Weekly Report)
- EID (Emerging Infectious Diseases)
- PCD (Preventing Chronic Disease)
We focus on PCD articles — a peer-reviewed open-access journal that publishes research on the causes, prevention, and control of chronic diseases such as diabetes, heart disease, obesity, and cancer. Each article is stored as HTML, preserving the original section structure (Abstract, Methods, Results, Discussion, etc.), which is critical for section-aware question generation.
The dataset is accessed via the Socrata Open Data API and downloaded locally. Each record contains the article’s full HTML body, publication metadata, and a unique identifier.
_URL_PCD = "https://data.cdc.gov/api/views/ut5n-bmc3/files/c0594869-ba74-4c26-bf54-b2dab3dff971?download=true&filename=pcd_2004-2023.zip"
HTML_ZIP_DIRECTORY="./cdc-corpus-data/zip"
if not Path(HTML_ZIP_DIRECTORY).exists():
print("No data.. downloading")
download_file(url=_URL_PCD,file_name="pcd.zip")Once downloaded, the zip is extracted to ./cdc-corpus-data/html-outputs/pcd/. The load_html_files utility reads each HTML file into a dictionary keyed by relative path.
data_dir = get_data_directory()
print(data_dir)
extract_zip_files()
target_dir = data_dir / "html-outputs/pcd"
if target_dir.exists():
data_html = load_html_files()/path/to/cdc-corpus-data
Loading HTML articles: 100%|██████████| 2914/2914 [00:58<00:00, 49.98file/s]
Loaded 2914 HTML articles
The full PCD corpus contains several thousand articles. For this walkthrough we sample the first 100 articles (itertools.islice(..., 100)) to keep generation and indexing time manageable. Scale SAMPLE_N and EVAL_SAMPLE_N at the end to run against the full corpus.
import random
import itertools
type(data_html)
#data_html_samples = random.choices(list(data_html.items()),k=50)
data_html_samples= {}
for k,v in itertools.islice(data_html.items(),100):
data_html_samples[k]= v1.2 Chunking: Semantic HTML Splitting
Raw HTML articles are parsed using a semantic-preserving HTML splitter that respects heading hierarchy (h1, h2, h3). This produces chunks that are naturally aligned with paper sections — each chunk corresponds to a coherent unit of scientific content rather than an arbitrary character window.
This is preferable to a generic RecursiveCharacterTextSplitter for structured scientific documents: it avoids splitting a Methods paragraph mid-sentence and prevents mixing content from unrelated sections (e.g., merging a Results paragraph with a Discussion paragraph). Preserving section boundaries is especially important for section-aware question generation in Part 2.
Each chunk carries metadata: doc_id (article identifier) and section (inferred from the heading hierarchy).
from langchain_core.documents import Document
from langchain_text_splitters import RecursiveCharacterTextSplitter, HTMLSectionSplitter
from langchain_text_splitters import HTMLSemanticPreservingSplitter
headers_to_split_on = [
("h1", "Title"), # Article titles (PCD, MMWR)
("h2", "Section"), # Main sections in PCD (Abstract, Methods, Results, Discussion)
("h3", "SubSection"), # Main sections in EID/MMWR, subsections in PCD
]
splitter = HTMLSemanticPreservingSplitter(
headers_to_split_on=headers_to_split_on,
separators=["\n\n", "\n", ". ", "! ", "? "],
max_chunk_size=3000,
chunk_overlap=350,
preserve_images=False,
preserve_videos=False,
elements_to_preserve=["ul", "ol"],
denylist_tags=["script", "style", "head"],
)
documents = []
for relative_path, content in data_html_samples.items():
#html_sections = text_splitter.split_text(content['html_content'].replace("\n", ""))
html_sections = splitter.split_text(content['html_content'])
doc_html_sections = []
for doc in html_sections:
metadata = doc.metadata
metadata.update({'doc_id':relative_path.split("/")[-1]})
new_doc = Document(page_content=doc.page_content,metadata = metadata)
doc_html_sections.append(new_doc)
documents.extend(doc_html_sections)1.3 Filtering Chunks by Size
Not all chunks are suitable for question generation. Very short chunks (e.g., figure captions, headers) lack enough information to generate meaningful questions. Very long chunks overwhelm the LLM context and produce low-quality, diffuse statements.
We keep only chunks between 2,000 and 10,000 characters — a range that captures complete paragraphs and multi-paragraph sections while excluding boilerplate text. These bounds are configurable via MIN_CHUNK_SIZE and MAX_CHUNK_SIZE in the config cell.
# --- Configuration ---
MIN_CHUNK_SIZE = 2000 # minimum chunk character length
MAX_CHUNK_SIZE = 10000 # maximum chunk character length
SAMPLE_N = 2 # contexts per section for question generationprint("Total Context:", len(documents))
# Remove chunks that are too small (likely navigation/headers)
filtered_documents = [
doc for doc in documents
if len(doc.page_content) >= MIN_CHUNK_SIZE
]
print(f"Total Context >= {MIN_CHUNK_SIZE} chars:", len(filtered_documents))
Total Context: 1963
Total Context >= 2000 chars: 705filtered_documents = [
doc for doc in filtered_documents
if len(doc.page_content) <= MAX_CHUNK_SIZE
]
print(f"Total Context <= {MAX_CHUNK_SIZE} chars:", len(filtered_documents))
Total Context <= 10000 chars: 7021.4 Filtering by Section
A key insight from the Know Your RAG paper is that different question types are best suited to different parts of a scientific article:
| Section | Best question type | Why |
|---|---|---|
| Results, Methods, Measures, Data Analysis | fact_single | Contain precise, verifiable claims: statistics, procedures, measurements |
| Discussion, Abstract, Conclusion, Implications | reasoning | Contain inferred conclusions, interpretations, and synthesized insights |
We filter chunks to retain only sections that will actually generate questions, discarding Introduction, Background, and References sections which are either too generic or too citation-heavy for useful RAG evaluation questions.
This section mapping is defined in reasoning_sections and fact_sections and drives all downstream generation logic.
# Prepare data for the DataFrame
data = []
for doc in filtered_documents:
row = {"page_content": doc.page_content}
row.update(doc.metadata) # Add all metadata key-value pairs
data.append(row)
# Create the Pandas DataFrame
df_filtered = pd.DataFrame(data)
df_filtered.head()df_filtered['merge_section'] = df_filtered['Section'].fillna(df_filtered['SubSection'])The keep list below is a broad allowlist that discards noise sections (References, footnotes, boilerplate). active_sections — defined next — is the narrower subset that actually drives question generation. Introduction and Background are retained in the DataFrame but are never sampled for question generation.
keep = ['Introduction',
'Background',
'Results',
'Discussion',
'Methods',
'Abstract',
'Implications for Public Health',
'Intervention Approach',
'Evaluation Methods',
'Implications for Public Health Practice',
'Purpose and Objectives',
'Outcomes',
'Highlights',
'Summary',
'Conclusion',
'Measures',
'Statistical analysis',
'Data collection',
'Analysis',
'Data analysis',
'Statistical analyses',
'Data sources',
'Intervention']
reasoning_sections = [
'Discussion',
'Abstract',
'Implications for Public Health',
'Implications for Public Health Practice',
'Conclusion',
'Summary',
]
fact_sections = [
'Results',
'Methods',
'Measures',
'Statistical analysis',
'Data collection',
'Analysis',
'Data analysis',
'Statistical analyses',
'Data sources',
'Evaluation Methods',
]
## filter null sections
df_filtered_clean = df_filtered[df_filtered['merge_section'].notna()]
df_filtered_section = df_filtered_clean[df_filtered_clean['merge_section'].isin(keep) ]
df_filtered_section.head()Part 2 — Evaluation Dataset Generation
2.1 Sampling Contexts
Generating questions for every chunk in the corpus would be expensive. Instead, we take a stratified sample of SAMPLE_N chunks per active section type. This ensures the evaluation dataset covers a variety of sections and topics rather than being dominated by whichever section is most common in the corpus.
The sample is reproducible (random_state=42) and respects section boundaries — each sampled chunk will produce questions of the appropriate type for its section.
active_sections = reasoning_sections + fact_sections
df_active = df_filtered_section[df_filtered_section['merge_section'].isin(active_sections)]
df_final_sample = (
df_active
.groupby('merge_section', group_keys=False)
.apply(lambda x: x.sample(n=min(SAMPLE_N, len(x)), random_state=42))
.reset_index(drop=True)
[['page_content', 'doc_id', 'merge_section']]
)
df_final_sample
2.2 Statement and Question Generation
The Know Your RAG framework generates questions indirectly: rather than asking an LLM to write a question directly from a passage, it first extracts intermediate statements and then generates a question for each statement. This two-step process produces more grounded, specific questions and makes critique easier (since you can verify the statement against the context independently of the question).
The pipeline for each passage or chunk is:
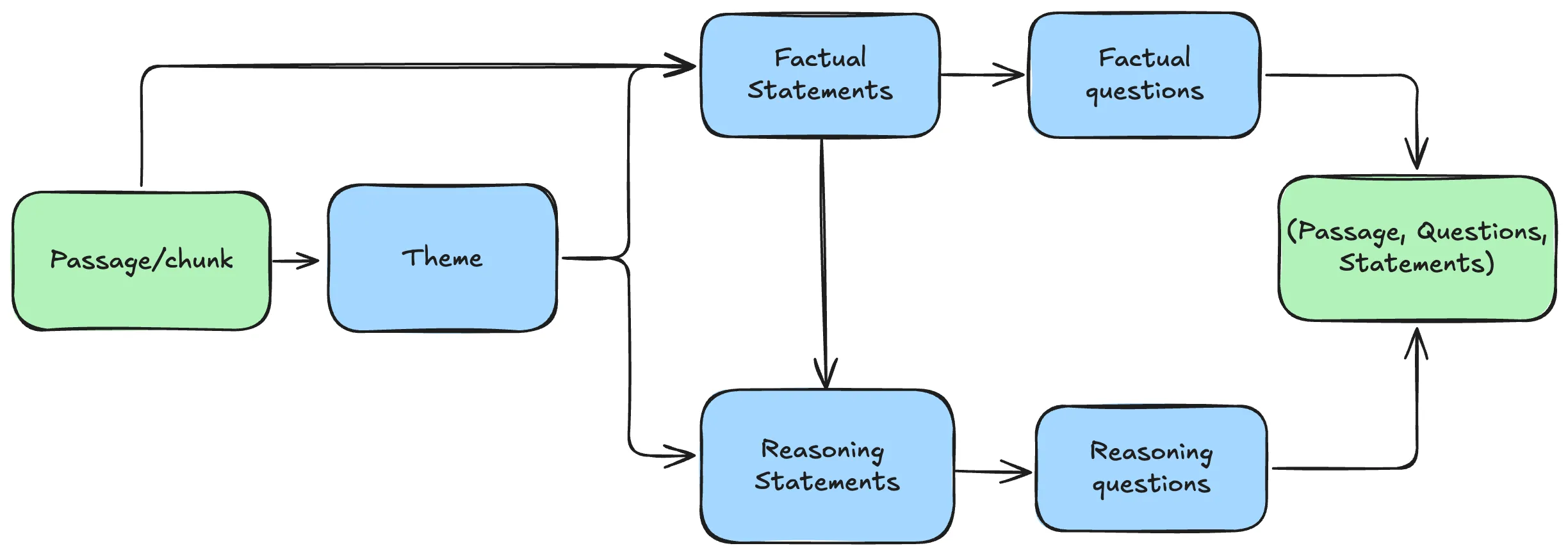
All generation functions accept a provider argument ('openai' or 'ollama') so you can swap the underlying model without changing the pipeline logic.
class Statements(BaseModel):
statements: List[str] = Field(default_factory=list, description="list of statements")
class Question(BaseModel):
question: str = Field(description="A self-contained question, with no references to any specific study, document, or author")def call_llm(provider: str, instructions: str, prompt: str, response_format=None):
"""
Call an LLM provider.
Args:
provider: 'openai' or 'ollama'
instructions: System prompt
prompt: User prompt
response_format: Pydantic model class for structured output (e.g. Statements, Question),
or None for plain text
Returns:
Parsed Pydantic object if response_format is set, otherwise a plain string.
"""
messages = [
{"role": "system", "content": instructions},
{"role": "user", "content": prompt},
]
if provider == 'ollama':
client = openai.OpenAI(
base_url="http://localhost:11434/v1",
api_key="ollama"
)
if response_format is not None:
response = chat(
model="llama3.2",
messages=messages,
format=response_format.model_json_schema()
)
return response_format.model_validate_json(response.message.content)
else:
response = client.chat.completions.create(model="llama3.2", messages=messages)
return response.choices[0].message.content
elif provider == 'openai':
client = openai.Client()
if response_format is not None:
response = client.beta.chat.completions.parse(
model="gpt-4o-mini",
messages=messages,
response_format=response_format
)
return response.choices[0].message.parsed
else:
response = client.chat.completions.create(model="gpt-4o-mini", messages=messages)
return response.choices[0].message.contentStep 1 — Theme Extraction
Before generating statements, we extract a short theme (2–5 words) from each chunk. The theme acts as a contextual anchor: it is injected into every downstream prompt to prevent generic statements like “the study found…” and instead encourage domain-specific phrasing like “diabetes prevalence among low-income adults…”.
This is especially important for epidemiological text, where different articles may describe similar methodologies in similar language. The theme makes generated questions distinguishable and retrievable.
def generate_theme(context: str, provider='openai'):
instructions = """ You are public health research assistant.
Your role is to read and understand paper passages
and extract the relevant and dominant theme in a few words."""
theme_prompt = f"In a few words, extract the main theme behind the following passage: {context}"
return call_llm(provider=provider, instructions=instructions,
prompt=theme_prompt, response_format=None)The helper functions below support all generation steps. generate_questions turns a statement into a standalone search query; is_standalone is a fast phrase-matching pre-filter that rejects questions containing context-referencing phrases before the expensive critique LLM is called.
def generate_questions(statement: str, theme="", provider='openai'):
instructions = f"""
A public health researcher is looking for information about the following topic:
{theme}
They want to find an answer to the following fact:
{statement}
Generate one question they would type into a research database to find this information.
Requirements:
- Self-contained: no references to "this study", "the document", "the context", "the authors", or "the paper"
- Use specific epidemiological domain terms (population group, condition, geographic scope,
disease, intervention,outcome, diagnosis,comparison,Phenomenon of Interest, Evaluation, Research Type)
- Style of a PubMed or Google Scholar search query
- The question should make sense by itself
"""
result = call_llm(provider=provider, instructions=instructions,
prompt="Generate one question as instructed.",
response_format=Question)
return result.question# Phrases that signal a question is anchored to a specific document/study context
# and will fail the stand_alone critique metric.
CONTEXT_REFERENCE_PHRASES = [
'in this study', 'in the study', 'in the document', 'in the context',
'in this analysis', 'in this paper', 'in this article', 'this research',
'the authors', 'the researchers', 'according to the study',
'as described in', 'as mentioned in', 'the participants in this',
]
def is_standalone(question: str) -> bool:
"""Return True if the question contains no implicit document/context references."""
q_lower = question.lower()
return not any(phrase in q_lower for phrase in CONTEXT_REFERENCE_PHRASES)deduplicate_questions and global_deduplicate (shown after) remove near-duplicate questions at the per-context and global levels respectively. This step is not in the original Know Your RAG paper — it was added here because LLMs reliably generate semantically similar questions when processing related chunks. In a corpus of epidemiological studies, many papers describe similar populations or methods in similar language, and the same question can surface independently from multiple passages. Leaving near-duplicates in the evaluation set inflates scores by testing the same thing repeatedly. While not a validated approach, TF-IDF cosine similarity is a practical and interpretable first pass for catching these before they reach the critique step: a strict threshold (0.80) is applied per-context to remove near-identical variants of the same question, and a relaxed threshold (0.85) is applied globally to remove cross-context duplicates without over-filtering genuinely similar but distinct questions.
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics.pairwise import cosine_similarity
def deduplicate_questions(questions: list, threshold: float = 0.80) -> list:
"""
Remove near-duplicate questions from a list of (statement, question) tuples.
Uses TF-IDF cosine similarity on the question text.
Preserves the first occurrence of each unique question.
Args:
questions: List of (statement, question) tuples
threshold: Similarity threshold above which a question is considered a duplicate.
0.80 = per-context (strict), 0.85 = global (relaxed)
Returns:
Deduplicated list.
"""
if len(questions) <= 1:
return questions
texts = [q for _, q in questions]
tfidf = TfidfVectorizer().fit_transform(texts)
sim_matrix = cosine_similarity(tfidf)
keep = []
dropped = set()
for i in range(len(texts)):
if i in dropped:
continue
keep.append(questions[i])
for j in range(i + 1, len(texts)):
if sim_matrix[i, j] >= threshold:
dropped.add(j)
return keep
def global_deduplicate(items: list, threshold: float = 0.85) -> list:
"""
Deduplicate (context, section, qtype, statement, question) tuples on question text
across all contexts.
"""
if len(items) <= 1:
return items
texts = [item[4] for item in items]
tfidf = TfidfVectorizer().fit_transform(texts)
sim = cosine_similarity(tfidf)
keep = []
dropped = set()
for i in range(len(items)):
if i in dropped:
continue
keep.append(items[i])
for j in range(i + 1, len(items)):
if sim[i, j] >= threshold:
dropped.add(j)
print(f'Global dedup: {len(items)} → {len(keep)} questions ({len(items) - len(keep)} removed)')
return keepStep 2 — Factual Statements
From each chunk, the LLM extracts up to three factual statements — precise, self-contained claims derived directly from the text. Each statement must:
- Contain a single unit of factual information
- Be understandable without the source document
- Include contextual information from the theme
- Not repeat information from other statements
These statements serve two purposes: they become the answers for fact_single questions, and they serve as inputs for the reasoning statement generation step.
def generate_fact_statements(context: str, theme="", provider='openai'):
instructions = f"""You are a public health researcher extracting precise factual claims from epidemiological literature.
Extract at most three factual statements from the following passage about: {theme}
Each statement must strictly follow these guidelines:
- State one specific, verifiable fact (a statistic, a measurement, a defined procedure, or a documented finding)
- Write it as a self-contained declarative sentence — no references to "this study", "the authors", "the paper", or "the analysis"
- Include specific epidemiological context: name the population group, geographic scope, time period, or condition where relevant
- Use precise language: prefer "adults with type 2 diabetes in low-income urban areas" over "study participants"
- Only include information explicitly present in the passage — do not infer or generalize
- Each statement must be independent of the others
- Maximize diversity: each statement must cover a different dimension of the passage.
Aim for variety across: statistical findings (rates, counts, proportions),
population characteristics (age, income, geography, condition),
methodological details (study design, measurement, data source),
and outcomes or interventions. Do not generate two statements about the same metric.
Bad example (avoid): "In this study, 34% of participants reported low physical activity."
Good example: "Among low-income urban adults aged 18–64, 34% reported engaging in no leisure-time physical activity."
Theme: {theme}
Passage: {context}"""
return call_llm(provider=provider, instructions=instructions,
prompt="Extract factual statements as instructed.",
response_format=Statements)Step 3 — Reasoning Statements
Reasoning statements are inferred conclusions drawn from multiple factual statements — they capture insights that are implied by the evidence but not directly stated. This mirrors how a public health analyst would synthesize research findings.
These statements become the answers for reasoning questions, which are used to evaluate whether a RAG system can retrieve and synthesize evidence across a chunk — not just surface a single fact. Reasoning questions are applied only to Discussion, Abstract, Conclusion, and Implications sections, where interpretive content is concentrated.
def generate_reasoning_statements(statements: List[str], theme="", provider='openai'):
instructions = f"""You are a public health analyst drawing evidence-based conclusions from epidemiological research findings.
From the factual statements below about {theme}, generate three reasoning conclusions.
A reasoning conclusion is an evidence-based inference that:
- Is supported by multiple statements together, but not directly stated in any single one
- Connects risk factors to population outcomes, identifies disparities, or infers public health implications
- Is written as a standalone, self-contained sentence — no references to "this study", "the research", "the authors", or "the findings above"
- Uses specific epidemiological language: name population groups, conditions, or contexts rather than saying "participants" or "the cohort"
- Avoids generic conclusions like "more research is needed" or "this highlights the importance of..."
Bad example (avoid): "This research suggests that interventions targeting study participants could be beneficial."
Good example: "Food insecurity among low-income adults with chronic disease is associated with higher rates of preventable hospitalization, suggesting that social determinants of health are stronger predictors of outcomes than clinical factors alone."
Each conclusion must be:
- Independent and non-overlapping with the other conclusions
- Specific enough that a public health professional could act on or evaluate it
- Understandable without access to the original passage
Theme: {theme}
Statements: {statements}
"""
return call_llm(provider=provider, instructions=instructions,
prompt="Generate reasoning conclusions as instructed.",
response_format=Statements)Step 4 — Question Generation
For each statement (factual or reasoning), the LLM generates one question framed as a public health database search query. The theme is injected to ensure domain-specific, precise phrasing. Critically, the prompt explicitly forbids references to “this study”, “the document”, or “the context” — the most common source of low stand_alone scores.
A lightweight pre-filter (is_standalone) catches obvious context-referencing phrases before the expensive critique LLM is called, saving cost and improving throughput.
The full generation loop iterates over all sampled chunks, applies the appropriate statement types per section, and collects all (statement, question) pairs into a results list for the critique step.
from tqdm.notebook import tqdm
results = []
skipped_questions = 0
duplicates_removed = 0
for index, row in tqdm(df_final_sample.iterrows(), total=len(df_final_sample),
desc='Generating questions', unit='context'):
content = row['page_content']
section = row['merge_section']
theme_content = generate_theme(context=content, provider='openai')
facts_statements = generate_fact_statements(context=content,
theme=theme_content, provider='openai')
# fact_single: only for fact_sections
fact_questions = []
if section in fact_sections:
for fact in tqdm(facts_statements.statements, desc=f' fact [{section}]',
unit='q', leave=False):
q = generate_questions(fact, theme_content, provider='openai')
if is_standalone(q):
fact_questions.append((fact, q))
else:
skipped_questions += 1
before = len(fact_questions)
fact_questions = deduplicate_questions(fact_questions, threshold=0.80)
duplicates_removed += before - len(fact_questions)
# reasoning: only for reasoning_sections
reason_questions = []
if section in reasoning_sections:
reason_statements = generate_reasoning_statements(statements=facts_statements.statements,
theme=theme_content, provider='openai')
for reason in tqdm(reason_statements.statements, desc=f' reasoning [{section}]',
unit='q', leave=False):
q = generate_questions(reason, theme_content, provider='openai')
if is_standalone(q):
reason_questions.append((reason, q))
else:
skipped_questions += 1
before = len(reason_questions)
reason_questions = deduplicate_questions(reason_questions, threshold=0.80)
duplicates_removed += before - len(reason_questions)
result = {
'context': content,
'section': section,
'theme': theme_content,
'fact_questions': fact_questions,
'reason_questions': reason_questions,
}
results.append(result)
total_questions = sum(len(r['fact_questions']) + len(r['reason_questions']) for r in results)
print(f'\nGeneration complete: {total_questions} questions kept')
print(f' Skipped (context-referencing): {skipped_questions}')
print(f' Removed (per-context duplicates): {duplicates_removed}')Generation complete: 45 questions kept
Skipped (context-referencing): 0
Removed (per-context duplicates): 0The generation loop produced 45 questions distributed across fact_single and reasoning types, one per statement extracted from each sampled chunk. Before these reach the evaluation set, each question is scored for quality by a separate critique LLM — the next step filters out questions that are vague, context-dependent, or poorly grounded.
2.3 Critique & Filter
Not all generated questions are suitable for evaluation. The Know Your RAG paper proposes using a separate critique LLM to score each (context, question, answer) triplet on multiple quality dimensions. We apply six metrics, each scored 1–5 by gpt-4o-mini:
| Metric | What it checks | Why it matters for public health RAG |
|---|---|---|
| qc_groundness | Is the question answerable from the context? | Core validity — prevents questions that require external knowledge not in the corpus |
| ac_groundness | Is the answer derivable from the context? | Critical for reasoning questions where LLM inference can drift beyond the source text |
| q_feasibility | Does the question carry enough signal for retrieval? | Ensures the retriever can find the relevant chunk — vague questions produce meaningless retrieval scores |
| stand_alone | Is the question self-contained? | Questions referencing “the study” or “this document” are unusable as retrieval queries |
| q_usefulness | Would this question be useful for RAG evaluation? | Filters trivial or unanswerable questions that inflate or deflate scores artificially |
| qa_tautology | Does the answer add information beyond the question? | Especially important for fact_single — answers that just restate the question measure nothing |
The two metrics from the original paper that we omit — c_usefulness and c_clarity — evaluate context quality, which is already enforced upstream by size filtering (2,000–10,000 characters) and section filtering.
class CritiqueAnswer(BaseModel):
evaluation: str = Field(description="Your rationale for the rating, as a brief and concise text")
rating: int = Field(description="your rating, as a number between 1 and 5")# 1 - q_to_c_groundedness
question_to_context_groundness = """
You will be given a context and a sentence that should be a question.
Your task is to provide a 'total rating' scoring how well one can answer the
given question unambiguously with the given context.
Give your answer on a scale of 1 to 5,
where 1 means that the question is not answerable at all given the context,
and 5 means that the question is clearly and unambiguously answerable with the context.
If the sentence provided is not actually a question, rate it as 1.
You MUST provide values for a brief and concise 'evaluation' and 'rating' in your answer.
Now here are the question and context.
Question: "{question}"
Context: "{context}"
"""
# 2 - ac_groundness
answer_to_context_groundness = """
You will be given a context, and a passage.
Your task is to provide a 'total rating' scoring how well the statements in the provided passage
can be inferred from the provided context.
Give your rating on a scale of 1 to 5, where 1 means that none of the
statements in the passage can be inferred from the provided context,
while 5 means that all of the statements in the passage can be unambiguously and entirely
obtained from the context.
You MUST provide values for a brief and concise 'evaluation' and 'rating' in your answer.
Now here are the context and statement.
Context: "{context}"
Passage: "{answer}"
"""
#3 - q_feasability
question_feasibility = """
You will be given a context and a question.
This context is extracted from a collection of passages,
and the question will be used to find it.
Your task is to provide a 'total rating' scoring how well
this context can be retrieved based on the specificity and pertinence of the question.
Give your answer on a scale of 1 to 5, where 1 means that it will be difficult
to find this context from this question due to lack of specificity or pertinence,
and 5 means that the context can clearly be found with information contained in the question.
You MUST provide values for a brief and concise 'evaluation' and 'rating' in your answer.
Now here are the question and context.
Question: "{question}"
Context: "{context}"
"""
# 4 stand_alone
stand_alone = """
You will be given a question. Your task is to provide a 'total rating' representing
how context-independent this question is.
Give your answer on a scale of 1 to 5,
where 1 means that the question depends on additional information to be understood,
and 5 means that the question makes sense by itself.
For instance, if the question refers to a particular setting using phrases like
'in the context', 'in the document', 'in this study', 'in this paper', 'the authors found',
or 'according to the article', the rating must be 1.
The questions can contain technical terms, acronyms, population groups, or epidemiological
concepts and still be a 5 — as long as a researcher could understand what is being asked
without access to any specific document.
For example:
- "What is the prevalence of type 2 diabetes among low-income adults in rural US counties?" → 5
(self-contained public health query, no document reference)
- "What percentage of participants in this study reported physical inactivity?" → 1
(refers to a specific study's participants)
- "What did the authors find about obesity rates?" → 1
(implicit reference to a specific document)
- "What chronic disease risk factors are associated with food insecurity in urban populations?" → 5
(standalone epidemiological question)
You MUST provide values for a brief and concise 'evaluation' and 'rating' in your answer.
Now here is the question.
Question: "{question}"
"""
#5 q_usefulness
question_usefullness = """
You will be given a question.
This question is to be used to find information in a collection of
documents.
Give your answer on a scale of 1 to 5, where 1 means that the question
is not useful at all, and 5 means that the question is extremely useful.
You MUST provide values for 'evaluation' and 'rating' in your answer.
Now here is the question.
Question: "{question}"
"""
#6 qa_tautology
question_answer_tautology = """
You will be given a question and passage its answer.
Your question is to judge whether this question and answer pair form a tautological exchange.
Give your answer on a scale of 1 to 5, where 1 means that the question and answer repeat the same information,
and 5 means that the answer is made of entirely new information.
You MUST provide values for 'evaluation' and 'rating' in your answer.
Now here are the question and its answer.
Question::: "{question}"
Answer::: "{answer}"
"""Each prompt above is rendered with the actual question, context, and answer values and sent to gpt-4o-mini as a structured-output call returning a CritiqueAnswer with rating and evaluation fields. generate_critiques orchestrates all six calls for a single (context, statement, question) triplet.
def call_critique_llm(instructions:str):
messages = [
{"role": "system", "content": instructions},
{
"role": "user",
"content": "Evaluate and generate rating as instructed",
}]
client = openai.Client()
response = client.beta.chat.completions.parse(
model="gpt-4o-mini",
messages=messages,
response_format=CritiqueAnswer)
return response.choices[0].message.parsed
def generate_critiques(context, statement, question):
critic = {
"qc_groundness": call_critique_llm(instructions=question_to_context_groundness.format(
context=context,
question=question)),
"ac_groundness": call_critique_llm(instructions=answer_to_context_groundness.format(
context=context,
answer=statement)),
"q_feasibility": call_critique_llm(instructions=question_feasibility.format(
question=question,
context=context)),
"stand_alone": call_critique_llm(instructions=stand_alone.format(
question=question)),
"q_usefulness": call_critique_llm(instructions=question_usefullness.format(
question=question)),
"qa_tautology": call_critique_llm(instructions=question_answer_tautology.format(
question=question,
answer=statement)),
}
return critic
%%time
from tqdm.notebook import tqdm
# Flatten all questions across contexts
all_questions_flat = [
(result['context'], result['section'], qtype, statement, question)
for result in results
for qtype, statement, question in (
[('fact_single', s, q) for s, q in result['fact_questions']]
+ [('reasoning', s, q) for s, q in result['reason_questions']]
)
]
# Global dedup across all contexts (catches cross-context near-duplicates)
all_questions = global_deduplicate(all_questions_flat, threshold=0.85)
data = []
for context, section, qtype, statement, question in tqdm(
all_questions,
desc='Generating critiques',
unit='question',
):
row = {
'context': context,
'section': section,
'question_type': qtype,
'question': question,
'answer': statement,
}
critics = generate_critiques(context, statement, question)
for key, val in critics.items():
row[key] = val.rating
data.append(row)
generated_questions = pd.DataFrame(data)
generated_questions.head(), len(generated_questions)
The critique scores are now attached to every question. We filter to retain only questions that pass all thresholds (groundedness ≥ 3, feasibility ≥ 3, standalone ≥ 3, usefulness ≥ 3) and save the result to generate_questions.csv. Persisting to disk lets you reload the filtered dataset in a fresh session without re-running the expensive generation and critique steps. Note that qa_tautology is tracked but not applied as a hard filter: reasoning statements are inferred conclusions that necessarily rephrase and synthesize the question’s topic, so they inherently score lower on this metric. A strict tautology gate would disproportionately drop valid reasoning questions.
generated_questions_filtered = generated_questions.loc[
(generated_questions["qc_groundness"] >= 3)
& (generated_questions["ac_groundness"] >= 3)
& (generated_questions["q_feasibility"] >= 3)
& (generated_questions["stand_alone"] >= 3) # re-enabled after prompt improvements
& (generated_questions["q_usefulness"] >= 3)
#& (generated_questions["qa_tautology"] >= 4)
]
generated_questions_filtered.to_csv("generate_questions.csv")
generated_questions_filtered.head()Reload from disk — safe to re-run from this cell without re-running Parts 2.1–2.3.
generated_questions = pd.read_csv('./generate_questions.csv',index_col=[0])We draw a random sample of EVAL_SAMPLE_N questions from the filtered dataset and convert it to a list of dicts (eval_ds) — the format expected by both RAG pipelines and RAGAS. Each record has context (the source passage), question, and answer (the original statement used as ground truth).
EVAL_SAMPLE_N =10 # number of questions to evaluate
df_eval = generated_questions[['context', 'question', 'answer']].sample(
n=EVAL_SAMPLE_N, random_state=42
).reset_index(drop=True)
df_eval
eval_ds = df_eval[['context', 'question', 'answer']].to_dict(orient='records')
eval_ds[:2]
The filtered evaluation dataset is now ready — a list of (context, question, answer) records that will serve as input to both RAG pipelines. Part 3 covers indexing the full PCD corpus into a vector store and running each architecture against this dataset to produce RAGAS scores.
Part 3 — RAG System Evaluation
3.1 Vector Store Setup
Before evaluating, we index the full PCD corpus into a ChromaDB vector store using sentence-transformers/allenai-specter embeddings — a model specifically trained on scientific paper titles and abstracts, making it well-suited for epidemiological literature.
The corpus is chunked using RecursiveCharacterTextSplitter (chunk size 1,000, overlap 200) to produce retrieval-optimized chunks that are smaller than the question-generation chunks used in Part 2. Separating indexing chunks from generation chunks is intentional: generation benefits from larger, section-complete contexts, while retrieval benefits from smaller, focused chunks.
Note: Two separate chunk sizes are used intentionally. Generation (Part 2) uses large section-aligned chunks (2,000–10,000 chars) so the LLM has full context for statement extraction. Retrieval (Part 3) uses smaller chunks (1,000 chars) so semantic search returns focused, specific passages. Mixing the two would degrade both generation quality and retrieval precision.
# Retrieval chunks: smaller than generation chunks for focused semantic matching
text_splitter = RecursiveCharacterTextSplitter(
chunk_size=1000,
chunk_overlap=200,
)
# Note: this chunking is handled internally by VectorStoreAbstract.chunking()The chunking() method applies this splitter internally — see vectorstore.py for the full implementation.
3.2 The Evaluation Process
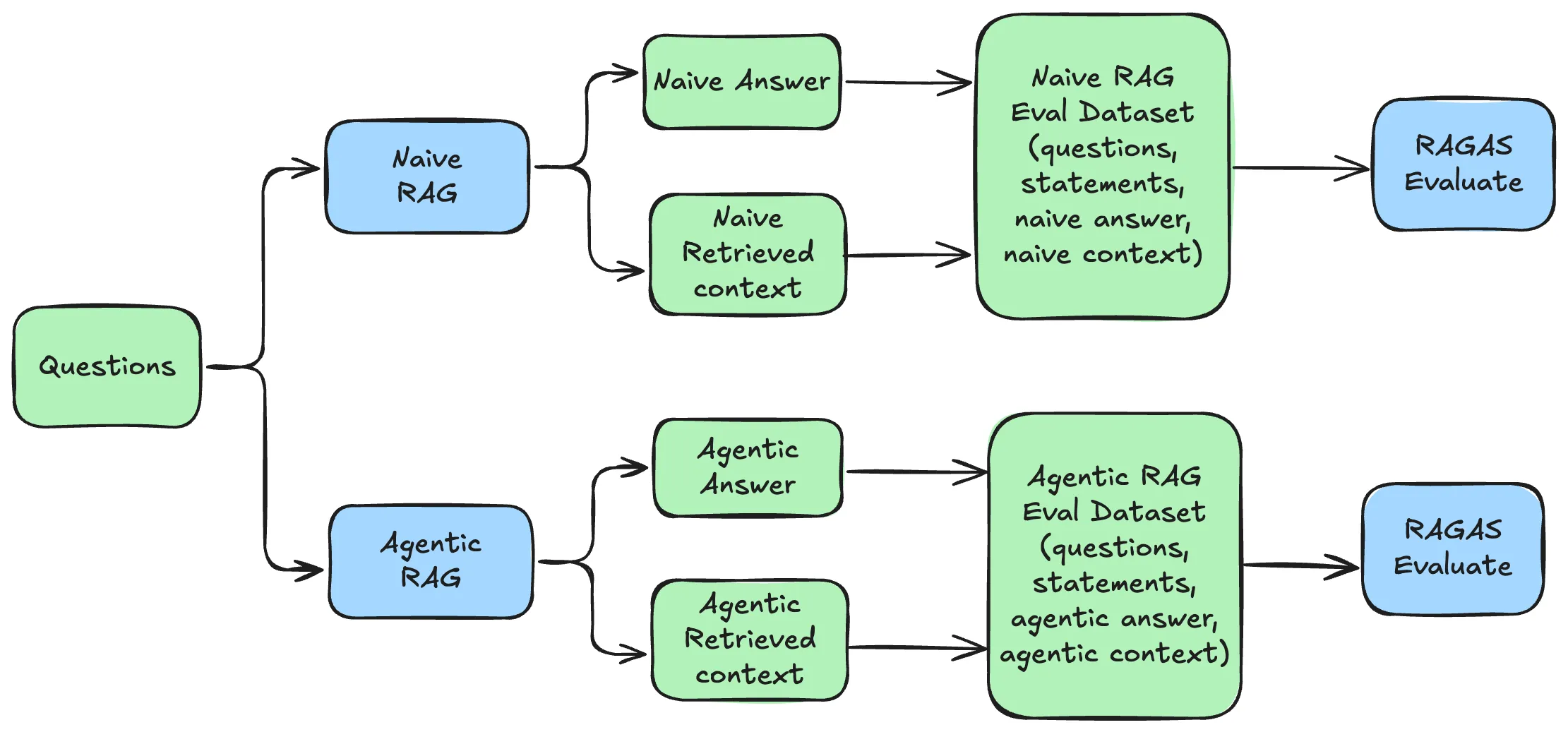
Each question from the evaluation dataset is passed to the RAG system, which retrieves relevant contexts and generates an answer. The resulting (question, answer, retrieved_contexts, reference) tuple is then scored by RAGAS. Running this loop independently for both architectures produces the per-question score tables compared in Part 4.
Sections 3.3 and 3.4 execute this loop on two different architectures — AgenticRAG first as the more complex system, NaiveRAG second as the baseline. Both produce CSV outputs (eval_results_agentic.csv, eval_results_naive.csv) that feed directly into the comparison table in Part 4.
3.3 AgenticRAG
The AgenticRAG system implements a multi-agent loop described in detail in Agentic RAG with OpenAI Agents SDK:
- Search agent — performs semantic search over the vector store for a given question or rephrased sub-question
- Evidence agent — scores each retrieved chunk for relevance (1–10) and summarizes the relevant content
- Answer agent — synthesizes the accumulated evidence into a final answer
The orchestrator loops until it has collected enough high-quality evidence (configurable via max_evidence_pieces and relevance_cutoff) or exhausted its search attempts (max_search_attempts). This architecture is designed to handle complex, multi-hop questions that require iterative retrieval — common in epidemiological research where a single question may require evidence from multiple sections of multiple papers.
from vectorstore import VectorStoreAbstract
CHROMA_PERSIST_DIRECTORY = './corpus-data/chroma_db'
# Convert all HTML-split documents to the format chunking() expects:
# {'content': text, 'id': doc_id}
abstracts = [
{'content': doc.page_content, 'id': doc.metadata.get('doc_id', '')}
for doc in documents
]
vector_store = VectorStoreAbstract(
abstracts=abstracts,
persist_directory=CHROMA_PERSIST_DIRECTORY,
recreate_index=True,
)
print(f'VectorStore initialized with {len(abstracts)} source documents')
Recreating existing index at ./corpus-data/chroma_db
Deleting existing collection...
Collection 'langchain' deleted successfully
New empty collection created
VectorStore initialized with 1963 source documents%%time
print('Chunking documents...')
documents_chunked = vector_store.chunking()
print(f'Created {len(documents_chunked)} chunks')
if vector_store.should_process_documents():
print('Indexing...')
vector_store.index_document(documents_chunked)
print(f'Done — {vector_store.get_document_count()} chunks stored')
else:
print(f'Using existing index ({vector_store.get_document_count()} chunks)')
RAGAS evaluation metrics require an LLM for scoring. We wrap gpt-4o-mini in a LangchainLLMWrapper for RAGAS compatibility. We use the legacy metric classes (ragas.metrics._answer_correctness, ragas.metrics._faithfulness) which are stable with the installed RAGAS version.
#%autoreload 0 # disable autoreload for ragas — causes harmless but noisy closure errors
import os
from dotenv import load_dotenv
from langchain_openai import ChatOpenAI
from ragas.llms.base import LangchainLLMWrapper
from ragas.dataset_schema import SingleTurnSample, EvaluationDataset
from ragas import evaluate
from ragas.metrics._answer_correctness import AnswerCorrectness
from ragas.metrics._faithfulness import Faithfulness
from ragas.run_config import RunConfig
load_dotenv(override=True)
run_config = RunConfig(timeout=300, max_retries=3)
evaluator_llm = LangchainLLMWrapper(ChatOpenAI(model='gpt-4o-mini'))
print('RAGAS ready')
from rag_agent import AgenticRAG, AgentConfig
config = AgentConfig(
collection_filter='pcd',
relevance_cutoff=8,
search_k=10,
max_evidence_pieces=5,
max_search_attempts=3,
mc_final_decision_only=False,
verbose=False,
)
agentic_rag = AgenticRAG(vector_store=vector_store, config=config)
# Smoke-test: ask a single question before running the full eval loop
test_question = eval_ds[0]['question']
answer, session = await agentic_rag.ask_question(test_question, max_turns=10)
print(f'Q: {test_question}')
print(f'A: {answer}')
Q: What characteristics and behaviors are associated with inadequate hydration among older adults in the United States?
A: Inadequate hydration among older adults in the United States is linked to several characteristics and behaviors:
1. **Demographic Factors**: Older individuals often experience decreased thirst sensation, resulting in lower drinking water intakes.
2. **Dietary Substitution**: Many older adults substitute water with calorically sweetened beverages, which detracts from proper hydration.
3. **Health Beliefs**: Individuals who believe their diet does not influence their health are more likely to drink less water. Those who show less concern for their health also tend to have lower hydration levels.
4. **Physical Activity**: Active older adults generally have higher water intake due to increased hydration needs from perspiration during exercise.
5. **Behavioral Patterns**: Lower water consumption often correlates with unhealthful habits, indicating that efforts to improve hydration should address these behavioral issues as well.
These factors underscore the complexity of hydration in older populations, highlighting the necessity for targeted public health interventions.from tqdm.notebook import tqdm
all_samples = []
for content in tqdm(eval_ds, desc='Evaluating', unit='question'):
answer, session = await agentic_rag.ask_question(
content['question'], max_turns=10
)
retrieved_contexts = [item['content'] for item in session.search_results]
sample = SingleTurnSample(
user_input=content['question'],
response=answer,
retrieved_contexts=retrieved_contexts,
reference=content['answer'],
)
all_samples.append(sample)
Evaluating: 0%| | 0/10 [00:00<?, ?question/s]print(f'\nRunning RAGAS on {len(all_samples)} samples...')
results_eval = evaluate(
dataset=EvaluationDataset(samples=all_samples),
metrics=[AnswerCorrectness(), Faithfulness()],
llm=evaluator_llm,
run_config=run_config,
)
results_df = results_eval.to_pandas()
results_df.to_csv('eval_results_agentic.csv', index=False)
print(results_df[['user_input', 'answer_correctness', 'faithfulness']])
results_evalRunning RAGAS on 10 samples...
Evaluating: 0%| | 0/20 [00:00<?, ?it/s]
user_input answer_correctness \
0 What characteristics and behaviors are associa... 0.349446
1 What is the correlation between fruit and vege... 0.550941
2 What is the impact of health insurance coverag... 0.434195
3 What is the prevalence of multiple chronic con... 0.309023
4 What is the prevalence of multiple chronic con... 0.392688
5 What are the factors associated with low drink... 0.223260
6 What are the associations between self-reporte... 0.201563
7 What factors contribute to low daily water int... 0.206767
8 What are the characteristics and behaviors ass... 0.403517
9 What is the prevalence of multiple chronic con... 0.215106
faithfulness
0 1.000000
1 0.312500
2 0.600000
3 1.000000
4 0.928571
5 0.882353
6 1.000000
7 0.333333
8 0.722222
9 0.250000
{'answer_correctness': 0.3287, 'faithfulness': 0.7029}3.4 NaiveRAG with Reranking
The NaiveRAG system represents the baseline single-pass approach:
- Retrieve — semantic search returns the top-k chunks from the vector store
- Rerank — ColBERT reranking re-scores all retrieved chunks and selects the top-N most relevant
- Generate — a single LLM call synthesizes the reranked chunks into an answer
This architecture has lower latency and cost than AgenticRAG and represents the standard production RAG pattern. Comparing it against AgenticRAG on the same evaluation dataset reveals whether the added complexity of an agentic loop improves answer quality for public health questions.
from rag_rerank import NaiveRAG, RerankConfig
rerank_config = RerankConfig(
top_k_retrieve=20,
top_rerank=3,
reranker_model='colbert',
verbose=False,
)
naive_rag = NaiveRAG(vector_store=vector_store, config=rerank_config)
print('NaiveRAG ready')NaiveRAG ready# Smoke-test: ask a single question before running the full eval loop
# output omitted for brevity — verify answer is non-empty before running the full eval loop
test_question = eval_ds[0]['question']
answer, status = naive_rag.ask_question(test_question)
print(f'Q: {test_question}')
print(f'A: {answer}')
print(f'Status: {status}')from tqdm.notebook import tqdm
naive_samples = []
for content in tqdm(eval_ds, desc='NaiveRAG Evaluating', unit='question'):
answer, status = naive_rag.ask_question(content['question'])
# Retrieve contexts separately so RAGAS faithfulness can check grounding
retrieved_chunks = vector_store.semantic_search(
query=content['question'],
k=rerank_config.top_k_retrieve,
)
retrieved_contexts = [c['content'] for c in retrieved_chunks]
sample = SingleTurnSample(
user_input=content['question'],
response=answer,
retrieved_contexts=retrieved_contexts,
reference=content['answer'],
)
naive_samples.append(sample)
print(f'\nRunning RAGAS on {len(naive_samples)} samples...')
results_naive = evaluate(
dataset=EvaluationDataset(samples=naive_samples),
metrics=[AnswerCorrectness(), Faithfulness()],
llm=evaluator_llm,
run_config=run_config,
)
results_naive_df = results_naive.to_pandas()
results_naive_df.to_csv('eval_results_naive.csv', index=False)
print(results_naive_df[['user_input', 'answer_correctness', 'faithfulness']])
results_naiveNaiveRAG Evaluating: 0%| | 0/10 [00:00<?, ?question/s]
Running RAGAS on 10 samples...
Evaluating: 0%| | 0/20 [00:00<?, ?it/s]
user_input answer_correctness \
0 What characteristics and behaviors are associa... 0.482706
1 What is the correlation between fruit and vege... 0.647744
2 What is the impact of health insurance coverag... 0.180394
3 What is the prevalence of multiple chronic con... 0.181669
4 What is the prevalence of multiple chronic con... 0.219457
5 What are the factors associated with low drink... 0.177618
6 What are the associations between self-reporte... 0.178577
7 What factors contribute to low daily water int... 0.215836
8 What are the characteristics and behaviors ass... 0.230669
9 What is the prevalence of multiple chronic con... 0.476626
faithfulness
0 1.000000
1 0.909091
2 0.000000
3 0.000000
4 1.000000
5 0.000000
6 0.000000
7 0.800000
8 1.000000
9 1.000000
{'answer_correctness': 0.2991, 'faithfulness': 0.5709}Part 4 — Comparison
4.1 RAGAS Evaluation Metrics
Both RAG systems are evaluated using RAGAS on the filtered evaluation dataset:
- Answer Correctness — measures semantic similarity between the generated answer and the reference answer (the original statement). Captures whether the system produces the right information.
- Faithfulness — measures whether the generated answer is grounded in the retrieved contexts. A high faithfulness score means the system’s answer can be traced back to retrieved evidence, indicating low hallucination risk.
For public health applications, faithfulness is particularly critical — a system that generates accurate-sounding answers not grounded in retrieved evidence is producing hallucinations, which is unacceptable in a clinical or policy context.
4.2 Results
Note: These scores are based on 10 questions — sufficient for a proof-of-concept but not for statistically robust conclusions. Increase
EVAL_SAMPLE_Nin the configuration cell to scale up before drawing any broader inferences.
import pandas as pd
metrics = ['answer_correctness', 'faithfulness']
agentic_scores = results_df[metrics].mean().rename('AgenticRAG')
naive_scores = results_naive_df[metrics].mean().rename('NaiveRAG')
comparison = pd.DataFrame([agentic_scores, naive_scores])
comparison.index.name = 'RAG'
comparison.columns = [m.replace('_', ' ').title() for m in metrics]
print('=== Mean scores across all questions ===')
print(comparison.to_string())
comparison=== Mean scores across all questions ===
Answer Correctness Faithfulness
RAG
AgenticRAG 0.328651 0.702898
NaiveRAG 0.299130 0.570909The table above is the core output of this pipeline: a side-by-side score comparison grounded in a domain-specific evaluation dataset built entirely from PCD literature. On this small sample, AgenticRAG produced higher scores on both metrics than NaiveRAG. The numbers reflect both the quality of the retrieval strategy and the quality of the evaluation questions — which is why investing in the generation pipeline matters.
Conclusion
This notebook demonstrated a complete, reproducible pipeline for building a domain-specific evaluation dataset and benchmarking RAG systems on epidemiological literature — from raw HTML articles to RAGAS scores. In this experiment, AgenticRAG produced higher answer correctness and faithfulness than NaiveRAG. This is consistent with the expectation that iterative evidence scoring helps on questions requiring synthesis across multiple passages — but 10 questions is not enough to draw general conclusions.
What we built
Starting from PCD articles in the CDC open corpus, we:
- Parsed and filtered documents using semantic HTML splitting, preserving section structure that is critical for question type assignment
- Generated a grounded evaluation dataset using the Know Your RAG framework — section-aware generation of
fact_singleandreasoningquestions, each traceable to a specific statement extracted from the source text - Enforced quality at every stage: diversity constraints in statement generation, standalone pre-filtering, six LLM-based critique metrics, and TF-IDF deduplication at both per-context and global levels
- Indexed the corpus into a ChromaDB vector store using
allenai-specterembeddings optimized for scientific text - Evaluated two RAG architectures — AgenticRAG (multi-agent loop) and NaiveRAG (retrieve → rerank → generate) — using RAGAS
AnswerCorrectnessandFaithfulness
Key takeaway
Generic evaluation datasets borrowed from other domains are a poor fit for public health RAG. Epidemiological text has domain-specific failure modes — study-scoped language, implicit population references, near-duplicate statistics across papers — that require targeted prompt engineering and filtering to produce meaningful evaluation questions. The multi-stage quality pipeline here (diversity constraints → standalone filter → critique → deduplication) addresses each of these systematically.
Limitations
Model dependency. All generation steps — statement extraction, question generation, critique scoring — and the final RAGAS evaluation used gpt-4o-mini. A stronger model (GPT-4o, Claude Sonnet) may produce higher-quality statements, better-calibrated critique scores, and different pass/fail decisions at the filter step. The pipeline architecture is model-agnostic, but the specific results here should not be assumed to generalize across model families or to smaller open-source models.
Critique threshold calibration. The filter threshold of ≥ 3 across all critique metrics is a pragmatic midpoint on a 1–5 scale — not derived from the original paper or validated against human judgment. A stricter threshold (≥ 4) produces fewer but higher-quality questions; a looser one (≥ 2) retains more questions but risks including weak or borderline ones that reduce evaluation signal. The right threshold depends on corpus size, acceptable evaluation set size, and downstream use. Calibrating these thresholds against human-annotated quality judgments is a natural next step before using this pipeline in production.
Next steps
- Scale up
SAMPLE_NandEVAL_SAMPLE_Nfor more statistically robust evaluation. - Extend to MMWR and EID articles to test generalization across CDC publication types.
- Add retrieval metrics (MRR, Recall@k, Context Precision) to diagnose whether failures are in retrieval.
- Add generation metrics (Response Relevancy, Topic Adherence) to diagnose failures in generation.
- Compare embedding models —
allenai-spectervs.all-MiniLM-L6-v2vs. a biomedical model likeBiomedBERT. - Extend and compare to other RAG frameworks such as GraphRAG, LightRAG, or RAPTOR.
- Additionally, such a domain-heavy tool must include review and input from subject matter experts.
References
- Lima, Rafael Teixeira de et al. “Know Your RAG: Dataset Taxonomy and Generation Strategies for Evaluating RAG Systems.” ArXiv abs/2411.19710 (2024). Paper
- Aymeric Roucher. RAG Evaluation. HuggingFace Cookbook. Notebook
- CDC. CDC Text Corpora for Learners — MMWR, EID, and PCD Articles. data.cdc.gov. Dataset